What is Applied Scientist
Applied Scientist is an autonomous agent that runs inside Jupyter. You hand it your baseline notebook and a research source (PDF, web URL, Kaggle link, GitHub/GitLab repo, or a plain-text idea), and it does the work a researcher would do by hand: read it, run your baseline, implement the new method, and produce a structured comparison. You supply the inputs, launch the run, and read the result.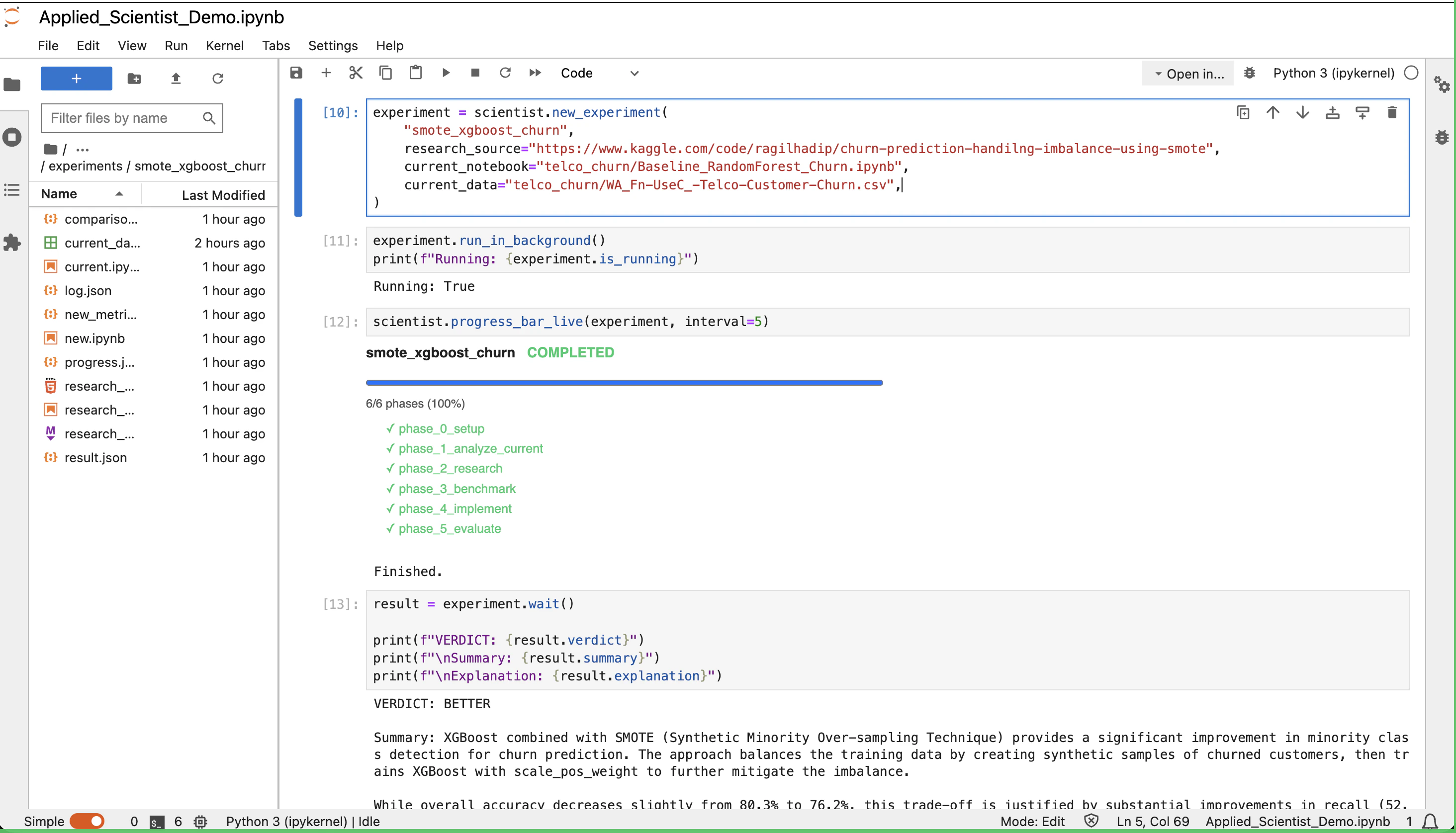
How It Works
A run moves through six fixed phases. Each phase has one job and hands off to the next.Phase 0 — Setup
Creates an isolated workspace and copies your notebook, data, and research source into it. The original files are never touched.
Phase 1 — Analyze Current
Reads your baseline notebook and documents the model, preprocessing, hyperparameters, and the metrics it reports.
Phase 2 — Research
Digests the research source: what the method does, what it improves, its requirements, and whether it’s compatible with your data.
Phase 3 — Benchmark
Locks in the metrics and baseline values that both sides will be measured on. Missing baseline metrics are flagged so the new run computes them too.
Phase 4 — Implement
Writes a new notebook implementing the method, using the same data, split, and seed as the baseline. Runs it end-to-end.
Cursor & Claude Code vs Upsonic Prebuilt Autonomous Agents
A question we hear a lot: why use this instead of just doing the same thing in Cursor or Claude Code? The short answer is that those are general coding copilots, and Applied Scientist is a purpose-built experiment runner. The table below shows where the two approaches diverge.| Dimension | Cursor & Claude Code | Upsonic Applied Scientist |
|---|---|---|
| Workspace | Runs in your working repo, shared with your editor | Fully isolated workspace folder per experiment |
| Output | Free-form chat and file edits | Structured ExperimentResult (verdict, comparison table, metrics) |
| Workflow | Assembled case by case in the chat | Pre-tested, well-designed pipeline |
| Environment | Outside the notebook | Runs directly inside Jupyter |
| Progress tracking | Scroll through chat transcript to guess where it is | Live progress bar driven by progress.json, plus last_logs(n) timeline |
Install
Requirements
You only need two things on disk.Baseline notebook
A working
.ipynb that trains your baseline model end-to-end. This is the reference every comparison is made against.Research source
Anything describing the method to try: PDF, Markdown/HTML, web URL, arXiv link, GitHub/GitLab/Bitbucket repo, Kaggle notebook or dataset page, or a free-form idea as plain text.
Running an Experiment
The example below is the demo shipped with Upsonic: a Random Forest baseline for telco customer churn, benchmarked against a Kaggle notebook that uses SMOTE + XGBoost to handle class imbalance.1. Create the agent
workspace is the root directory the agent is allowed to work in. Every experiment lives in its own folder inside it.
2. Define the experiment
research_source is polymorphic — pass any of these and the agent figures out how to materialize it:
- Local files — PDF, Markdown, HTML,
.ipynb, plain text - Web URLs — blog posts, arXiv pages, documentation
- Code hosts — GitHub, GitLab, or Bitbucket repository URLs
- Kaggle — notebook or dataset pages
- Free-form idea — a plain string describing what to try
| Parameter | Purpose |
|---|---|
name (positional) | Folder name and registry key |
research_source | Anything from the list above |
current_notebook | Path to your baseline notebook |
current_data | Optional. Data path or a short loader description. Inferred from the notebook when omitted. |
experiments_directory | Optional. Defaults to ./experiments inside the workspace. |
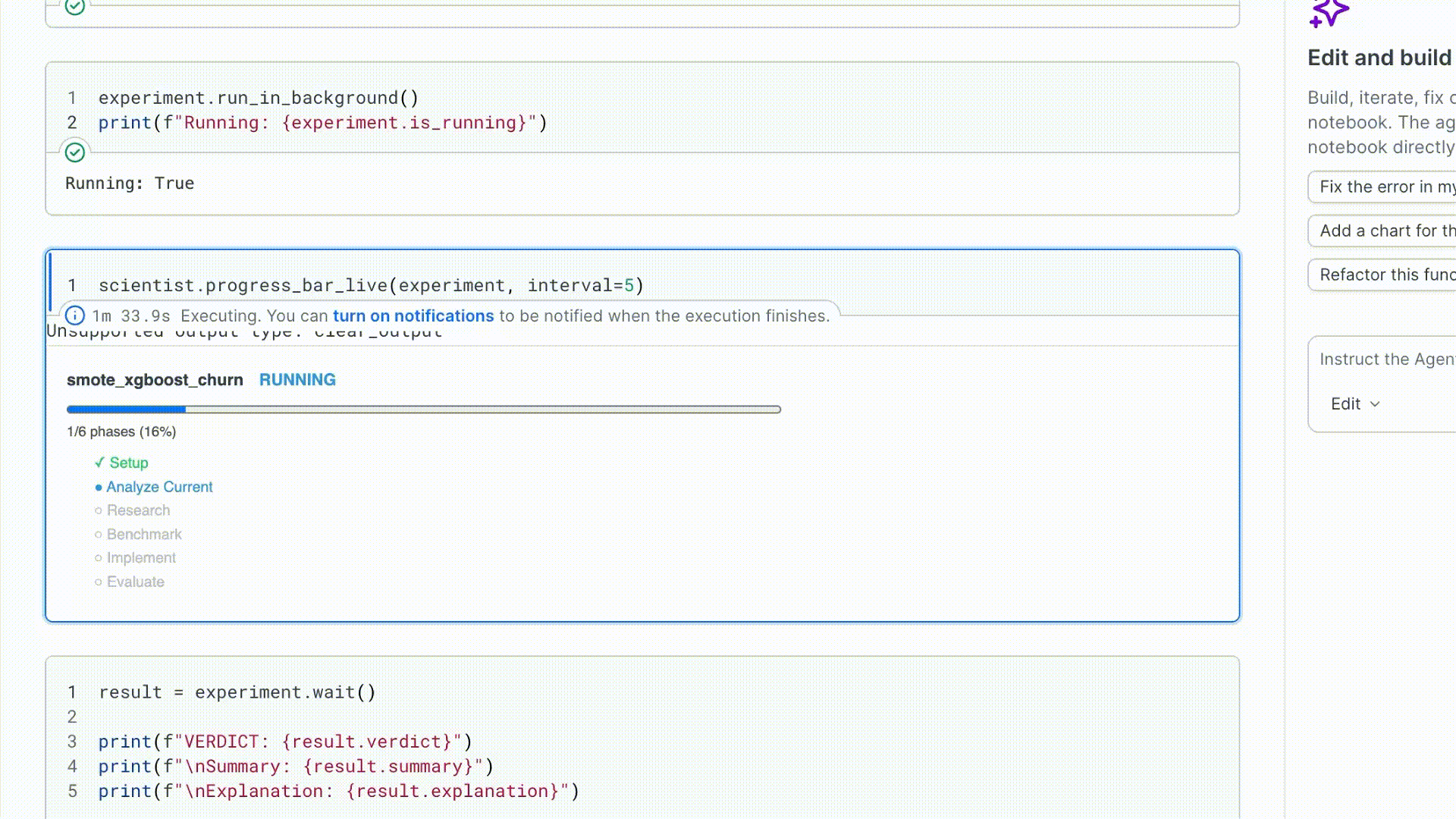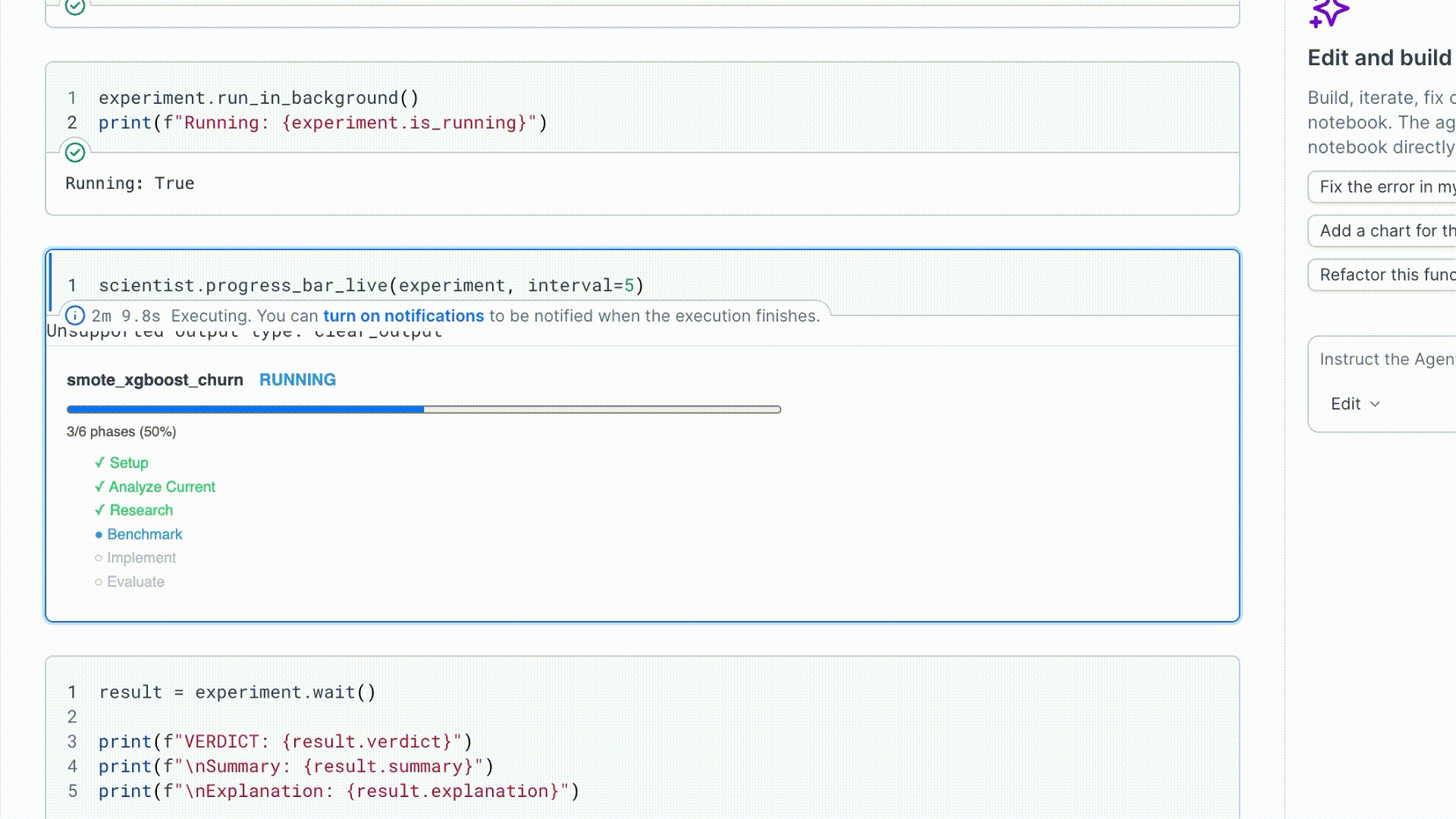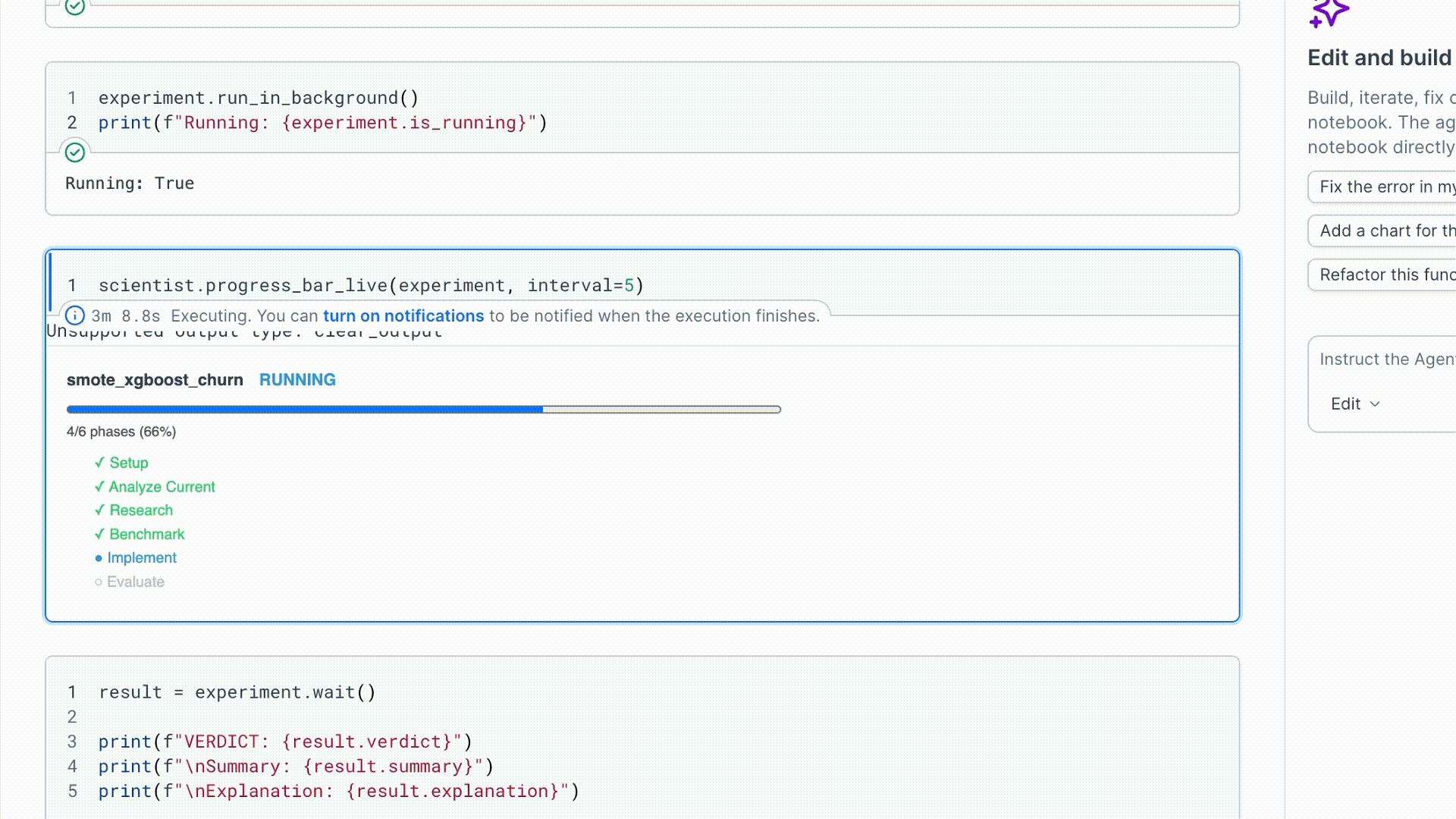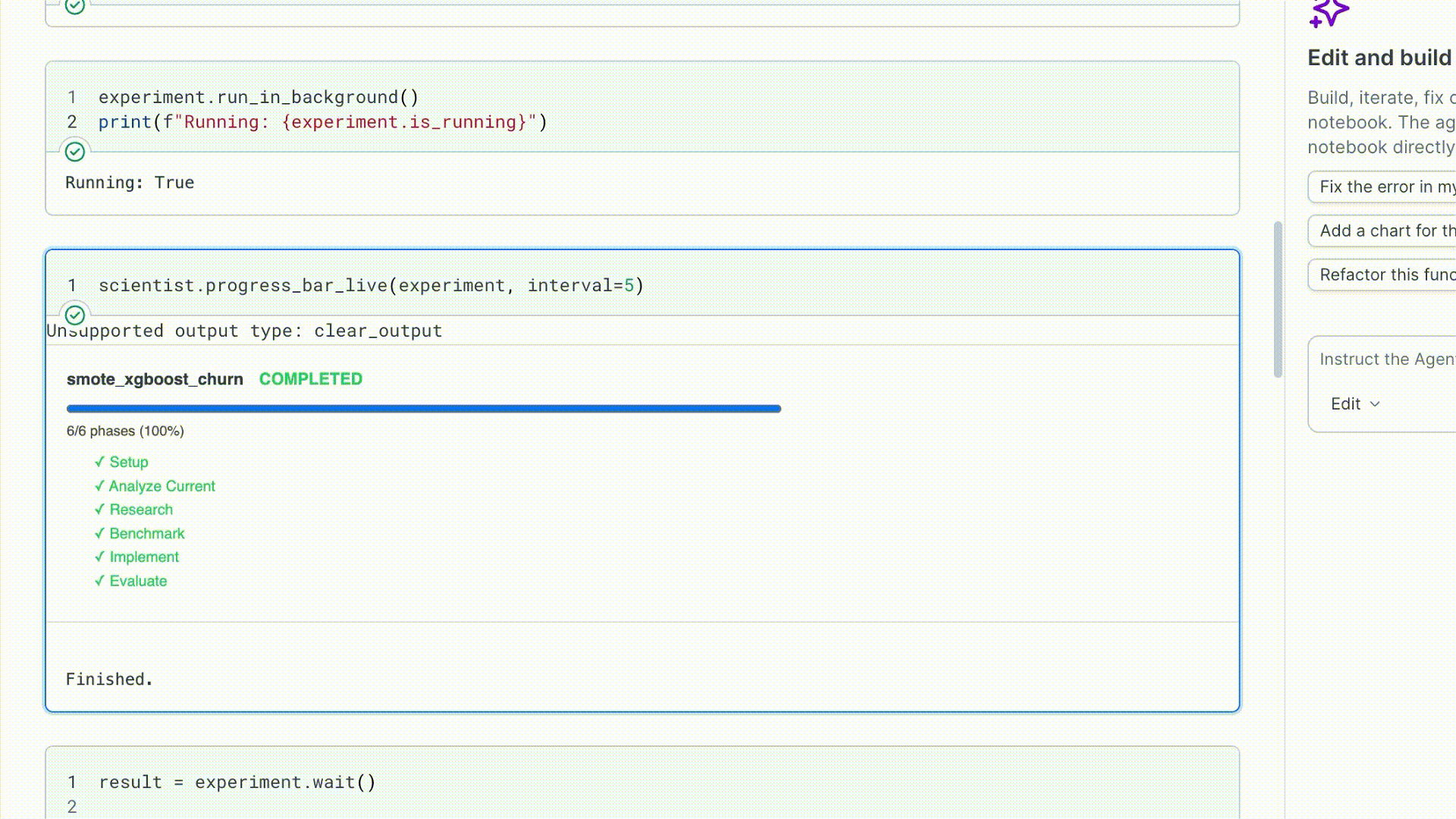
3. Run and watch
run_in_background() starts the run in a daemon thread and returns immediately.
| Attribute | What it tells you |
|---|---|
experiment.is_running | True while the thread is alive |
experiment.is_done | True once finished (success or error) |
experiment.error | The exception if the run raised, else None |
experiment.stop() to cooperatively cancel.
4. Wait for the result
wait() blocks until the run finishes and re-raises any exception it produced. For this demo run, it returns:
| Attribute | Value |
|---|---|
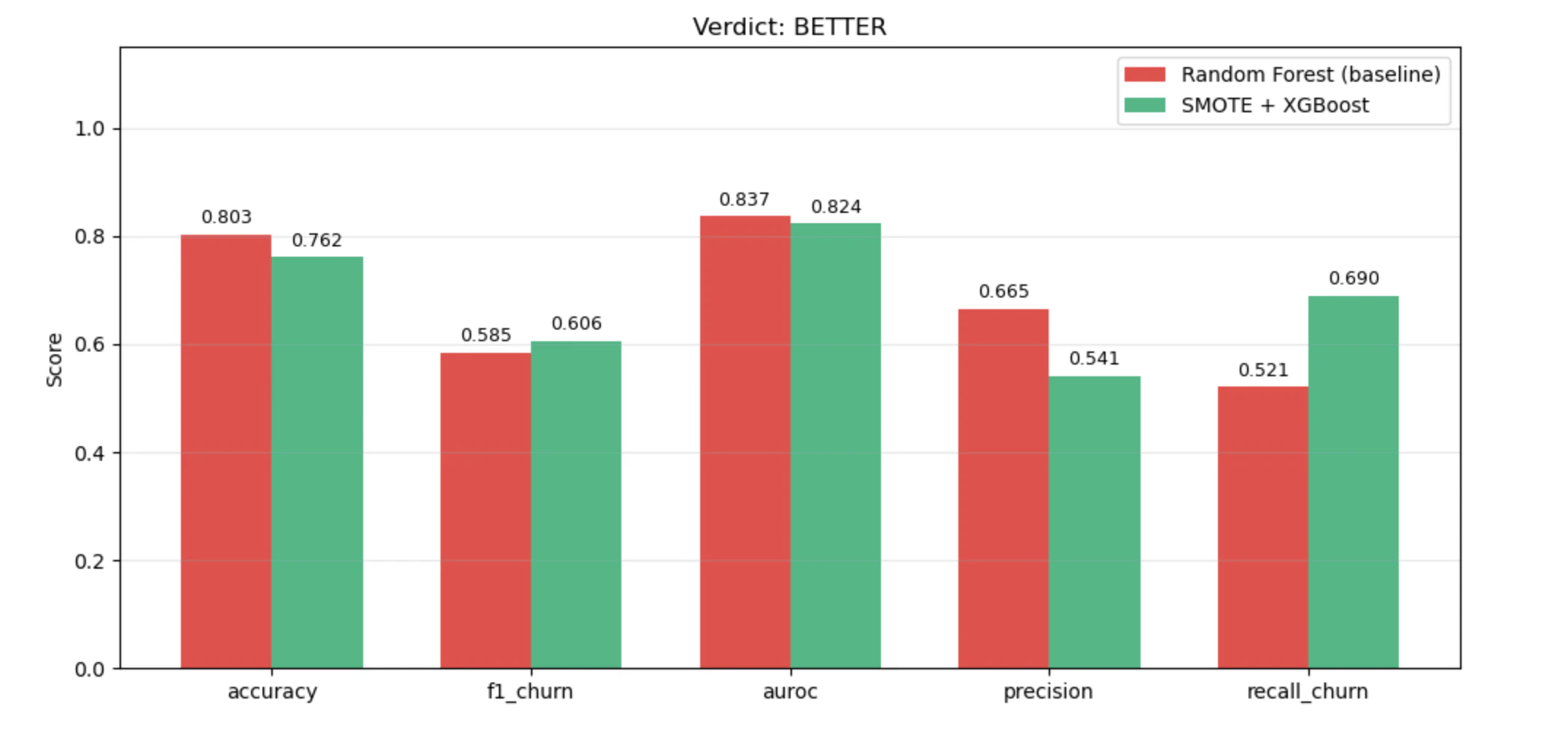
result.verdict | 'BETTER' | 'WORSE' | 'INCONCLUSIVE' | 'FAILED' |
result.summary | What the new method is and how it differs from the baseline |
result.explanation | Why this verdict was reached, referencing concrete numbers |
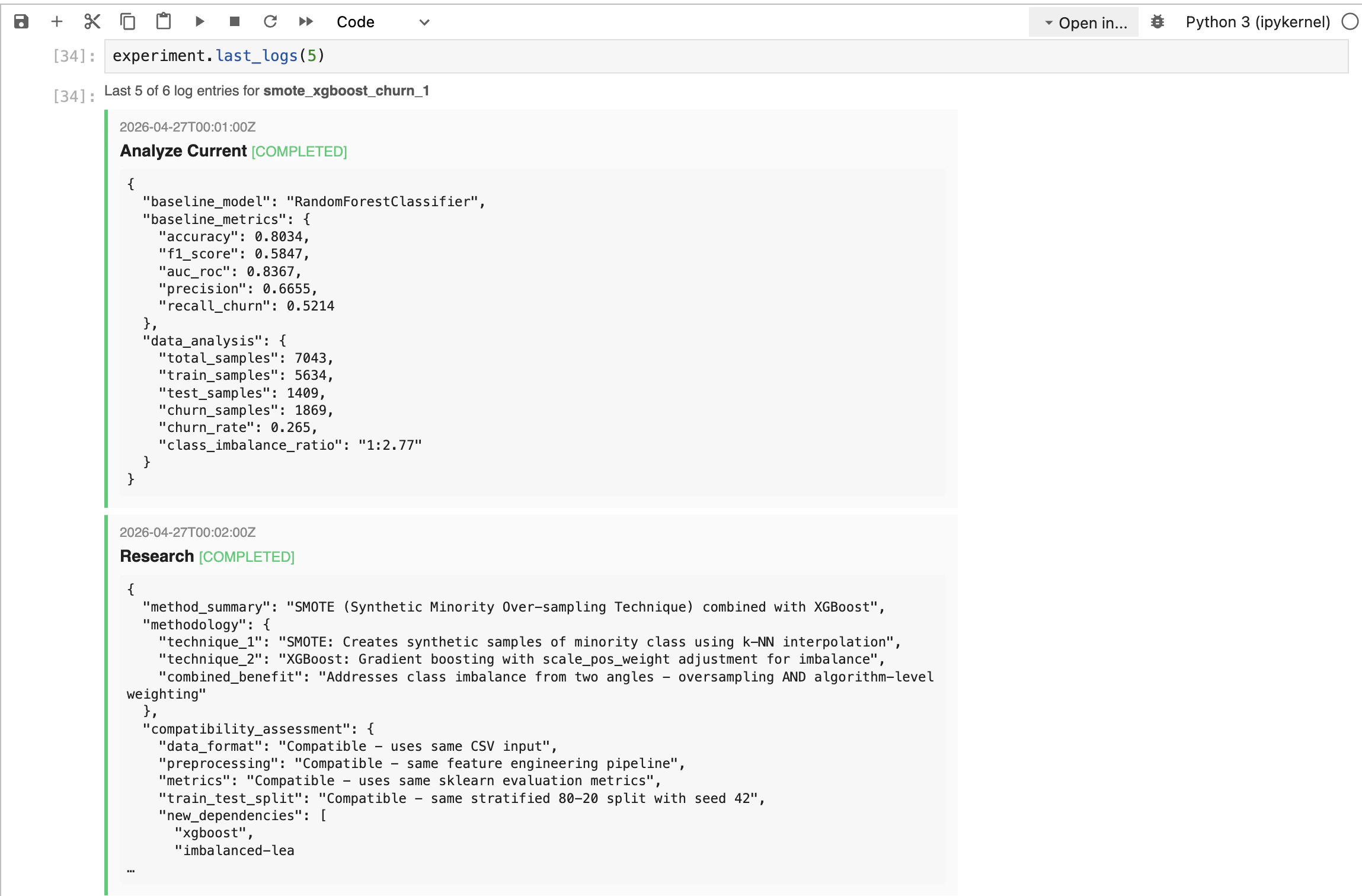
5. Inspect the comparison
result.table is a list of metric dicts. Drop it into a DataFrame to see the side-by-side:
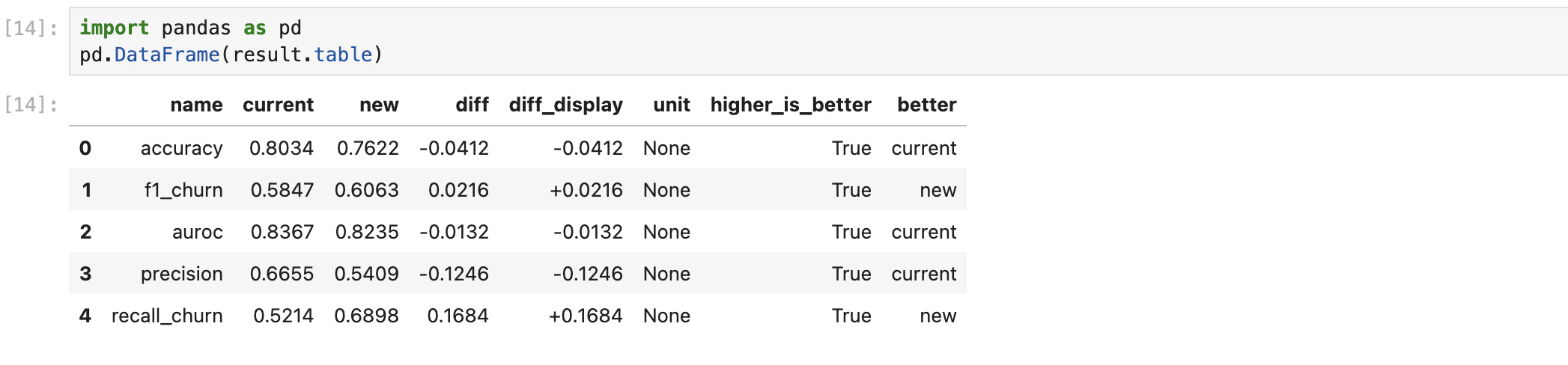
| Field | Meaning |
|---|---|
name | Metric name (e.g. accuracy, f1, auroc) |
current / new | Baseline and new-method values |
diff / diff_display | Raw difference and a human-friendly version |
unit | Unit of the metric |
higher_is_better | Whether larger is better |
better | Which side won on this metric (current or new) |
Need the raw artifacts?
result.record exposes log.json, progress.json, and registry metadata for the run.Managing Experiments
Every experiment is recorded inexperiments.json. The registry is re-read from disk on every call, so it always reflects current state.

name, date, status, verdict, baseline_model, new_method, paper, and path.
API Reference
The full demo notebook for this agent lives in the Upsonic repo under prebuilt_autonomous_agents.

